Forschungsgruppe Meier-Abt -Auf Englisch-
How protein abundance and protein structure determine the evolution of blood cancers
Our research assesses how a genotype via the proteotype leads to a phenotype in hematologic malignancies and cancer predisposition syndromes. A special focus is placed on the role of protein abundance and protein structural changes for the predisposition and disease evolution of hematologic neoplasms.
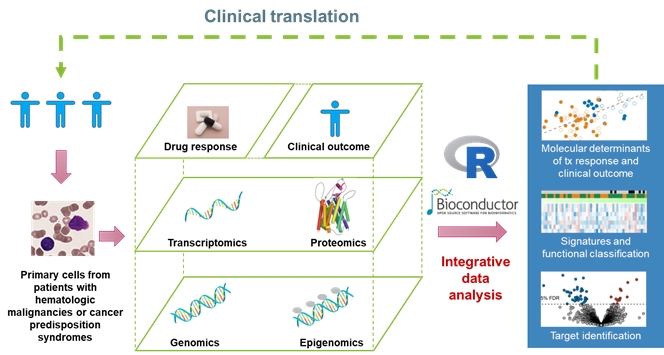
Figure: Multi-layered approach with focus on protein abundance and structure (proteotype) and its relation to genotype and clinical parameters.
Figure adapted from Meier-Abt F. Leading Opinions Hämatologie & Onkologie 2022 and from a graph by J. Lu. Protein image taken from Meier-Abt F. et al. J Membr Biol 2005.
Protein function determines cell functionality and development of disease. Protein abundance and structural changes are at the core of many signaling pathways and likely contribute to the evolution of hematologic malignancies. Targeted drugs rely on protein abundance and structural changes for activity. We recently generated protein abundance landscapes of different hematologic malignancies, including chronic lymphocytic leukemia (CLL) and the myeloproliferative neoplasm polycythemia vera (PV). We integrated the protein abundance data with genomic, transcriptomic, drug response and clinical outcome data for the same patients. A modification of the proteomic technique allows us to investigate protein structures in hematologic cancers. We will use this data to 1) assemble the protein abundance-protein structure-genotype landscape of hematologic malignancies, 2) link changes in protein structure to important genetic disease drivers, 3) assess the role of protein structure for drug response and clinical outcome.
We hypothesize that DNA damage repair pathways are important for disease evolution and emergence of treatment resistance. The role of protein abundance and protein structure for response and resistance to targeted drugs will be assessed in patients with somatic defects in DNA damage repair. We further hypothesize that protein structures associated with DNA damage repair defects are important for cancer-predispositions. The role of protein abundance and protein structures for risks of disease manifestation (penetrance) will be assessed in predisposed individuals with germline mutations in DNA damage repair genes.
To address these research aims, we employ recently developed mass spectrometry workflows in patient samples collected at the Dept. of Medical Oncology and Hematology (University Hospital Zurich) and the Institute of Medical Genetics (University of Zurich). Protein abundance and structural data is integrated with genomic and clinical data for the same individuals. Identified protein candidates are tested for causality and potential clinical translation in ex vivo systems.
Our work contributes to the integration of proteogenomic information into improved diagnostic and treatment strategies. It serves as proof-of-principle for other biological systems.